“This is one of those rare times where you can really tie everything back to a eureka moment,” study lead Dan Jacobson, of the US Department of Energy’s Oak Ridge National Laboratory (ORNL) in Tennessee, said in a news release.
“I was looking at data, and I suddenly saw some very distinct patterns happening in the pathways of the renin-angiotensin and bradykinin systems. That led us to do a deep dive of the gene families of the blood pressure regulatory system.”
The release described a phenomenon called a cytokine storm: a severe reaction that occurs when the human body produces more cytokines, “a variety of small proteins that help regulate the immune system,” than necessary.
Cytokine storms have been blamed for the mysteriously higher death rate seen in some illnesses, such as influenza, among younger people with stronger immune systems, which effectively overreact to the illness.
Early COVID-19 research was carried out to determine whether these cytokine storms were causing a variety of symptoms prevalent in those with COVID-19, including the loss of smell, skin lesions known as “COVID toes” and heart damage.
However, Jacobson believes this is related to the bradykinin pathway, which, along with the renin-angiotensin system, “regulates blood pressure and fluid balance in the body” and could actually help explain the different ways in which the novel coronavirus attacks the human body.
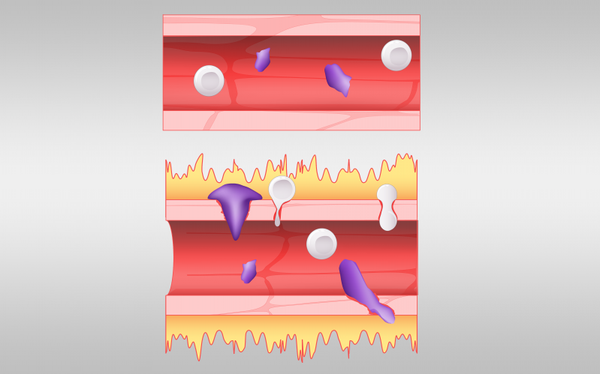
“We believe that when you take the inhibition at the top of this pathway off, you end up with an out-of-control cascade that leads to an opening up of the blood vessels, causing them to leak,” Jacobson said. “If that happens in the lung, that’s not good. Immune cells that are normally contained in the blood vessels flood into the surrounding infected tissue, causing inflammation.”
“Jacobson and his colleagues required the power of Summit to run 2.5 billion correlation calculations that helped them understand the normal regulatory circuits and relationships for the genes of interest,” according to the news release.
The results, published in peer-reviewed, open-access scientific journal eLife, revealed that genes associated with the bradykinin system “appear to be excessively ‘turned on’ in the lung fluid cells of those with the virus,” according to the news release.
If Jacobson and his colleagues’ hypothesis proves correct, then at least 10 existing drugs could be used to treat the pathways disrupted by the contagious disease, “but large-scale clinical trials are needed to determine whether they might be effective at treating COVID-19,” highlighted the release.